Shutterstock.com
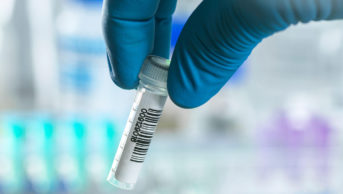
Many people in Australia can walk into their local pharmacy to request a pharmacogenomics (PGx) test, which, with some level of certainty, predicts how their body will respond to over 150 different drugs (see Box 1). Requiring only an inner cheek swab, each test would cost less than £100 to deliver in the UK. Within one week, the patient, community pharmacist and the patient’s GP receive a clinical report, which allows the prescriber to select an individualised treatment and dose to improve the chance of therapeutic success and minimise the likelihood of adverse drug events (see Table 1). Patients are required to take the test only once in a lifetime, and the results can be used to inform the safety and efficacy of all future prescribing decisions for the patient.
Box 1: Drugs for which pharmacogenomics testing can be used to provide prescribing guidance
- Angiotensin receptor blockers
- Anti-anginals
- Anti-arrhythmics
- Anticholinergics
- Anticholinesterases
- Anticoagulants
- Antidepressants (selective serotonin reuptake inhibitors, serotonin noradrenaline reuptake inhibitors, tricyclics, and others)
- Antidiabetics
- Anti-emetics
- Anti-epileptics
- Antifungals
- Antihistamines
- Antiplatelets
- Antipsychotics
- Antitussives
- Benzodiazepines
- Betablockers
- Calcineurin inhibitors
- Hypnotics
- Non-steroidal anti-inflammatory drugs
- Opioid analgesics
- Proton pump inhibitors
- Psychostimulants
- Statins
Similar multiple-drug testing services are already available in the United States[1]
,[2]
and Canada[3]
. In Norway, healthcare professionals can order a PGx test for any patient to test for any of the 150 drugs, at any time, as part of routine practice. In these countries, pharmacists are increasingly seen as central to the implementation of PGx testing[4]
.
As part of its strategy for personalised medicine, NHS England aims for whole-genome sequencing to become standard practice for specific conditions by 2020, along with “tailored, optimised and more effective therapies” by 2025[5]
. Despite leaps in gene technology and the increasing popularity of home genetics tests, such as 23andMe®, PGx testing is still rarely used in UK healthcare. The delay in the technology’s implementation in the UK presents an opportunity for pharmacy to place itself at PGx testing’s core for medicines optimisation.
| Personalised interpretation and recommendations | ||
| Medication | Interpretation | Recommendation |
| ♦ Codeine/ paracetamol | CYP2D6 — poor Greatly reduced metabolism of codeine into its active metabolite morphine. High likelihood of an inadequate analgesic response to codeine. | The Clinical Pharmacogenetics Implementation Consortium (CPIC) strongly recommends avoidance of codeine use owing to the lack of efficacy. CPIC states that tramadol (and oxycodone to a lesser extent) are not suitable alternatives. Examples of opioids not metabolised via CYP2D6 (and therefore not affected by this genetic variation) include morphine and buprenorphine. |
| ♦ Simvastatin | SLCO1B1 — low transporter function
| Based on the SLCO1B1 genotype, CPIC guidelines strongly recommend to be alert to the increased risk of myopathy, to consider low-dose therapy (20mg daily) and, if this is inappropriate, to consider an alternative statin. If using simvastatin, creatine kinase measurement may be considered. No genotype-guided dosing recommendation based on the CYP3A4 genotype is available. |
| â—Š Esomeprazole | CYP2C19 — extensive (normal) This genotype predicts normal metabolism of esomeprazole. However, this rate of metabolism leads to more rapid clearance of the drug and potentially to an incomplete clinical response in conditions such as oesophagitis and Helicobacter pylori infection. The effect of this genotype in predicting a reduced proton pump inhibitor response is less pronounced with esomeprazole than with several other drugs in this class (omeprazole, lansoprazole, pantoprazole). | If response is inadequate, consider:
|
| †Clopidogrel | CYP2C19 — extensive (normal) Normal formation of clopidogrel’s active metabolite is predicted. | CPIC guidelines strongly recommend use of the label-recommended dosage if clopidogrel is being prescribed for acute coronary syndrome with percutaneous coronary intervention. |
Source: Clinical Pharmacogenetics Implementation Consortium. Available at: https://cpicpgx.org/ (accessed May 2018); ♦ = Major prescribing considerations; ◊ = Minor prescribing considerations; †= Usual prescribing considerations | ||
Predicting a patient’s response to drugs
For many years, pharmacists have confidently estimated renal function to optimise therapy. We have known which drugs are metabolised through the liver and how they can interact with each other via this route, but we have largely based dosing information of these drugs on assumed population characteristics — a dose range likely to be safe and effective for most people. For patients with high metabolic rates, the maximum licensed dose may not be sufficient, and in patients with low metabolic rates, a low dose may actually have toxic effects. But now, by testing eight different cytochrome P450 genes, we can predict how patients will respond to different treatments and personalise therapy accordingly.
PGx: the evidence
Drugs are therapeutically effective in 20–60% of patients[6]
, and 38–75% of patients show no response to initial treatment[7]
. An individualised approach to prescribing, or ‘personalised medicine’, could increase the speed and likelihood of therapeutic response and reduce the possibility of adverse drug events[8]
,[9]
. The one-off cost of the test could be justified by a reduction in NHS costs as a result of fewer adverse events, as well as fewer visits to the prescriber.
Evidence is emerging to support both the effectiveness and cost-effectiveness of PGx testing at the point of medication initiation within individual drug groups such as warfarin[10]
,[11]
,[12]
, clopidogrel[13]
,[14]
, and antidepressants[15]
,[16]
. Most research on the subject to date has been performed in the United States[12]
,[15]
, with some in Hong Kong[12]
, China[14]
and the UK[11]
. A large-scale EU study is underway to investigate if PGx testing’s approach “results in a better outcome for patients and is cost effective”[17]
; the results will be used to develop strategies to make PGx testing available to every European citizen.
There is limited evidence to suggest how multiple-drug PGx testing should be used or implemented within medicines optimisation strategies. In a study of 112 care home residents in the United States, medications were reviewed using PGx test results; 15 patients had drug changes recommended based purely on their result, and many other recommendations were supported by the PGx test[18]
. However, patient benefit derived from this service was not captured or reported.
Another trial conducted in the United States compared pharmacist-led medication reviews and PGx testing with usual care (no medication review) through a randomised controlled trial in 110 patients, and found significant reductions in emergency department visits and hospitalisation[19]
. The study’s design did not allow the effect of PGx testing to be differentiated from that of the pharmacist review; consequently, the additional benefit and value of PGx testing in medicines optimisation remains unknown.
Pharmacy’s central role
PGx testing is not yet routine practice in the UK, so patients may need reassurance of the test’s purpose, how their DNA will be used and how their personal information will be stored and protected. Following clinical interpretation of the results, patients should also be made aware of the importance of the result for both current and future prescribing decisions. The pharmacist can support both the implementation of the test and patient discussion on receipt of the result.
PGx in primary care
One of the most obvious places for a PGx test is the point of medication initiation and it may be wise to undertake the test before making drug and dosing decisions. This could be carried out in community pharmacy, as in the Australian model, or in general practice by either the practice-based pharmacist or GP. On receipt of the test result, the relevant prescriber can discuss it with the patient and can select the most appropriate treatment and dose.
In an alternative model, the GP initiates therapy empirically and concurrently undertakes PGx testing, but asks the patient to discuss the result with their community pharmacist once it has been received. If there is no need for a change, the pharmacist can reassure the patient and explain what the results mean for future therapy. Otherwise, the pharmacist could refer the patient back for treatment alteration, which they would recommend to the GP. This could be incorporated into the new medicines service with appropriate training. Pharmacist prescribers could make the changes themselves.
Patients experiencing polypharmacy — concurrent prescription of five or more medicines[20]
— are obvious contenders for PGx testing and it could be incorporated into the medicines use review process. Suspicion of non-adherence could be roused if no side effects have been reported despite a high dose and the patient is a slow metaboliser. Similarly, in patients who are fast metabolisers and taking low doses, the treatment could be having no effect and may be suitable for deprescribing or dose changes.
Residents in care homes are also likely to benefit from PGx testing; the pharmacokinetics of many types of medicine commonly prescribed in this setting are predictable by the test (see Box 1).
PGx in secondary care
There could also be a place for PGx tests in hospital pre-admission clinics where patients are likely to be prescribed any of the testable drugs listed in Box 1, and it may also become commonplace in surgery, where testable drugs such as analgesia, warfarin and proton pump inhibitors are frequently used. Hospital pharmacists could use the results to make recommendations regarding both current and future therapy.
It may also be appropriate to request PGx testing when medicines are initiated before a patient is discharged. The patient should later discuss the result with their community pharmacist.
Becoming the linchpin
PGx testing is becoming routine practice in many countries and it is an area in which pharmacists can operate without eroding the roles of other healthcare professionals. Pharmacists have scientific training and are central to the medicines optimisation agenda, and implementation of PGx testing in the UK would be a natural extension of the pharmacy role. Its incorporation into the redesigned new medicine service and medicine use review processes could significantly benefit patients.
PGx testing and practical training for its implementation should be included in undergraduate pharmacy education in the UK, and, as in Australia, training should be readily available to pharmacists already in practice. Like calculation of renal function, PGx testing could eventually become second nature to pharmacists, so the profession must start to plan for its centrality to the practice now.
David Wright is professor of pharmacy practice at the University of East Anglia School of Pharmacy. Dhiren Bhatt is executive consultant at myDNA, Life Europe, Middlesex, UK. Correspondence to: d.j.wright@uea.ac.uk or dhiren.bhatt@mydna.life
References
[1] National Institute for Healthcare Excellence. Furosemide. Available at: https://bnf.nice.org.uk/drug/furosemide.html (accessed June 2018)
[2] OneOme. OneOme RightMed test: pharmacogenomic testing for medication drug response. Available at: https://oneome.com/ (accessed June 2018)
[3] Gray LJ, Taub NA, Khunti K et al. The Leicester Risk Assessment score for detecting undiagnosed type 2 diabetes and impaired glucose regulation for use in a multiethnic UK setting. Diabet Med 2010;27(8):887–895. doi: 10.1111/j.1464-5491.2010.03037.x
[4] Owusu-Obeng A, Weitzel KW, Hatton RC et al. Emerging roles for pharmacists in clinical implementation of pharmacogenomics. Pharmacotherapy 2014;34(10):1102–1112. doi: 10.1002/phar.1481
[5] NHS England. Improving outcomes through personalised medicine: Working at the cutting edge of science to improve patients’ lives. 2016. Available at: https://www.england.nhs.uk/wp-content/uploads/2016/09/improving-outcomes-personalised-medicine.pdf (accessed Jule 2018)
[6] Wilkinson GR. Drug metabolism and variability among patients in drug response. N Engl J Med 2005;352(21):2211–2221. doi: 10.1056/NEJMra032424
[7] Spear BB, Heath-Chiozzi M & Huff J. Clinical application of pharmacogenetics. Trends Mol Med 2001;7(5):201–204. PMID: 11325631
[8] Streetman DS. Emergence and evolution of pharmacogenetics and pharmacogenomics in clinical pharmacy over the past 40 years. Annals Pharmacother 2007;41(12):2038–2041. doi: 10.1345/aph.1K273
[9] Ahmed S, Zhou Z, Zhou J et al. Pharmacogenomics of drug metabolizing enzymes and transporters: relevance to precision medicine. Genomics Proteomics Bioinformatics 2016;14(5):298Â313. doi: 10.1016/j.gpb.2016.03.008
[10] Kim DJ, Kim HS, Oh M et al. Cost-effectiveness of genotype-guided warfarin dosing in patients with mechanical heart valve replacement under the fee-for-service system. Appl Health Econ Health Policy 2017;15(5):657–667. doi: 10.1007/s40258-017-0317-y
[11] Verhoef TI, Redekop WK, Langenskiold S et al. Cost-effectiveness of pharmacogenetic-guided dosing of warfarin in the United Kingdom and Sweden. Pharmacogenomics J 2016;16(5):478–484. doi: 10.1038/tpj.2016.41
[12] Gor D, Kim K, Chumnumwat S et al. Cost-effectiveness of a novel pharmacist guided warfarin pharmacogenetic service. Value Health 2015;18(7):A390. doi: 10.1016/j.jval.2015.09.866
[13] Wang Y, Yan BP, Liew D et al. Cost-effectiveness of cytochrome P450 2C19 *2 genotype-guided selection of clopidogrel or ticagrelor in Chinese patients with acute coronary syndrome. Pharmacogenomics J 2017;18(1):113–120. doi: 10.1038/tpj.2016.94
[14] Jiang M, You JH. Cost-effectiveness analysis of personalized antiplatelet therapy in patients with acute coronary syndrome. Pharmacogenomics 2016;17(7):701–713. doi: 10.2217/pgs-2016-0008
[15] Hornberger J, Li Q, Quinn B. Cost-effectiveness of combinatorial pharmacogenomic testing for treatment-resistant major depressive disorder patients. Am J Manag Care 2015;21(6):e357–365. PMID: 26247576
[16] Rosenblat JD, Lee Y & McIntyre RS. Does pharmacogenomic testing improve clinical outcomes for major depressive disorder? A systematic review of clinical trials and cost-effectiveness studies. J Clin Psychiatry 2017;78(6):720–729. doi: 10.4088/JCP.15r10583
[17] European Commission Community Research and Development Information Service. Ubiquitous pharmacogenomics (U-PGx): Making actionable pharmacogenomic data and effective treatment optimization accessible to every European citizen. 2016. Available at: https://cordis.europa.eu/project/rcn/199754_en.html (accessed June 2018)
[18] Sugarman EA, Cullors A, Centeno J et al. Contribution of pharmacogenetic testing to modeled medication change recommendations in a long-term care population with polypharmacy. Drugs Aging 2016;33(12):929–936. doi: 10.1007/s40266-016-0412-z
[19] Elliott LS, Henderson JC, Neradilek MB et al. Clinical impact of pharmacogenetic profiling with a clinical decision support tool in polypharmacy home health patients: A prospective pilot randomized controlled trial. PloS One 2017;12(2):e0170905. doi: 10.1371/journal.pone.0170905
[20] Masnoon N, Shakib S, Kalisch-Ellett L et al. What is polypharmacy? A systematic review of definitions. BMC Geriatr 2017;17(1):230. doi: 10.1186/s12