
istockphoto.com
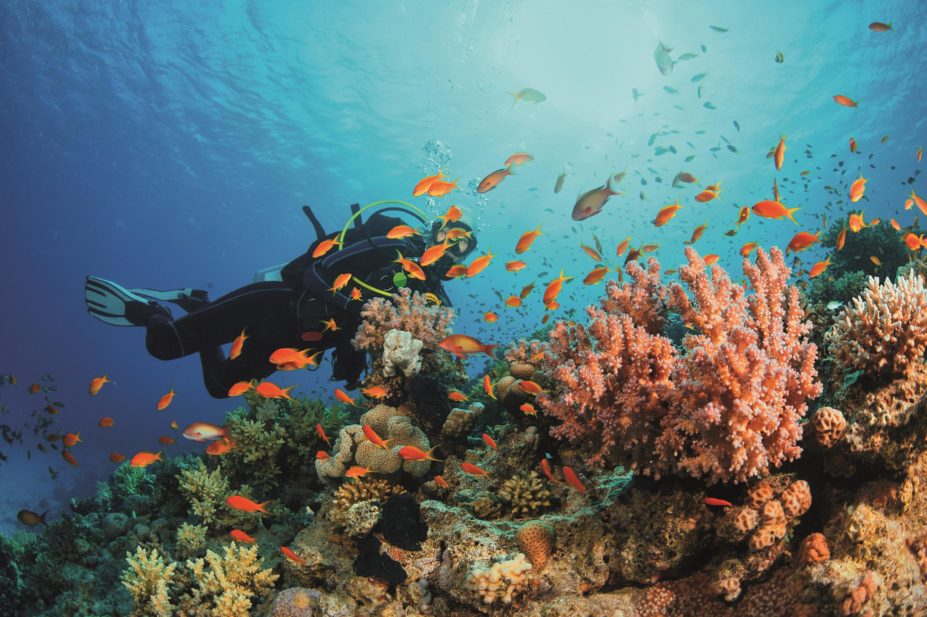
Nestled between the Arabian Peninsula and Africa is the Red Sea, snaking up from the Indian Ocean. It is home to thousands of species of marine life, and is a popular destination for scuba divers, thanks to its coral reefs and brightly coloured fish. But not all the visitors are there purely to enjoy the spectacle. Armed with plastic bags and knives, a group of researchers dive to a depth of 20 metres in their quest to discover new drugs.
The researchers, from the bacterial natural products laboratory at the Swiss Federal Institute of Technology in Zurich, along with collaborators from the department of zoology at Tel Aviv University are diving to collect sea sponges.
In January 2014, Joern Piel, professor of microbiology and head of the Swiss Federal Institute of Technology, published a paper that demonstrated what scientists had suspected for a long time — that the rich collection of compounds found in sponges are often the work of the symbiotic bacteria that share their home[1]
. “We found that one bacterium was producing dozens of different compounds and belongs to potentially a whole new phylum,” he says.
The aim of Piel’s research is to find new sources of biologically active molecules that could be used for drug development. “If you look at the diversity of current antibiotics that are in clinical use then you can see that most of them come from microorganisms. Bacteria are a rich source of bioactive products.” Piel is particularly interested in the drug discovery potential of uncultivated bacteria. But this could go far beyond antibiotics — natural products are used in a wide range of areas of medicine.
There is a huge diversity of compounds just waiting to be discovered
Scientists estimate that only 1% of the world’s microorganisms have been grown in the laboratory.[2]
“There is a huge diversity of compounds just waiting to be discovered,” says Piel.
Until relatively recently, the natural world was the only source of medicines. There is documented evidence that as long ago as 2,600 BC more than 1,000 species of plant were used to treat a range of ailments, and many of those are still in use today.[3]
Only in the past century has modern technology enabled chemists to create molecules of their own. There were hopes that this would lead to a torrent of new drugs, but it turns out that chemists are no rivals for nature.
So now scientists are finding new and efficient ways to hunt for drugs from nature — except this time they are searching primarily for microbes, rather than medicinal plants. Some scientists are using cutting-edge techniques to force these tiny life forms into manufacturing chemicals, which naturally are only produced under the most specific of circumstances. Others are literally going to the ends of the Earth to find microbes that have never previously been discovered.
Chemical failure
The 1950s and ’60s were the heyday of drug discovery from natural products and gave us many of the antibiotics and chemotherapies available today. However, the advent of personal computers in the 1980s changed everything. “Drug companies then had the capability to do very fast screening of molecules, and they quickly ran out of molecules to screen,” says David Newman, recently retired chief of the natural-products branch at the National Cancer Institute, based in Bethesda, Maryland. Drug screening leapt from a rate of about 100 compounds per week to more than 1,000 per week, he says. And since then it has only got faster, he adds, with GlaxoSmithKline (GSK) reportedly able to screen a quarter-of-a-million molecules per week.
When pharmaceutical companies ran out of molecules to screen, they used combinatorial chemistry to create libraries of thousands of molecules. This technique took off in the early 1990s, with pharmaceutical companies hoping it would revolutionise drug discovery. “They made molecules without anything to guide them, creating totally random libraries,” says Newman. At the same time, many companies closed their natural-products divisions, deciding that they didn’t fit in with their drug development timelines.
Yet despite the high hopes of the drug companies, these libraries of random molecules produced very few drugs. Since 1981, of all the new drugs that have been brought to market, only one, sorafenib, is the product of a random combinatorial library. By the end of the 1990s, people realised that they were not getting anywhere.
These compounds are so different from what a chemist can think of, it’s incalculable
“Making something from nothing has failed miserably,” says Newman. Mother Nature has had 3 billion years to refine her chemistry, he continues. “These compounds are so different from what a chemist can think of, it’s incalculable.”
Between 1981 and 2010, around 64% of the small-molecule drugs brought to market for cancer, infections and type 2 diabetes were based on natural products. Researchers showed that by using natural-product structures — which were not flat and had chiral centres — they could produce bioactive compounds using ‘combichem’ techniques but on a small scale (100 to 1,000, not 100,000 to >1,000,000 compounds). The other 36% were synthetic but not random[4]
.
Nature’s complexity is exemplified by the discovery of eribulin, which was brought to market by Eisai in 2010 to treat metastatic breast cancer. Eribulin is derived from halichondrin B, a natural product of the marine sponge[5]
. Piel says that scientists found that only about half of the original molecule was needed for bioactivity. “The original product has a very complicated structure and would need more sponges than exist on Earth to produce enough of it,” he says. So Eisai and collaborators managed to chemically synthesise the bioactive half, but even this is the “the most complex chemical synthesis of a drug that I know of”, says Piel.
Natural-products research is very big in academia. It could be a new golden age for natural products
Pharmaceutical companies still pick up natural products that have been discovered and bring them to market, but most companies no longer have their own dedicated natural-products divisions. Swiss company Novartis is a notable exception.
Academics, however, are more interested than ever in natural products. Piel is proof of this enthusiasm and he is far from alone. “Natural-products research is very big in academia. It could be a new golden age for natural products,” he says.
Exploiting microbes
Part of the reason for the resurgence in academic research is that genome sequencing has become much cheaper and faster. Historically, research was limited to molecules that could be produced by plants, or by bacteria that could be grown in the laboratory. There was a high rate of rediscovery of the same molecule, which put many researchers off.
“Chemists want new structures and a lot of medicinal plant compounds have already been characterised,” says Camila Esguerra, group leader at the Biotechnology Centre of Oslo, Norway. In comparison, microbes are a huge untapped source of molecules. But getting them to give up their secrets is not easy.
Researchers at NovoBiotic Pharmaceuticals — a spin-off company from Northeastern University in Boston, Massachusetts — have met this challenge by developing a system that can cultivate bacteria that were previously impossible to grow in the laboratory. “We really do not know what is absent in synthetic media that is needed for the majority of bacteria to be cultivated,” says Losee Ling, vice-president of research and development at NovoBiotic. However, using NovoBiotic’s approach, it is no longer necessary to know what the magic ingredient is, as the bacteria are grown in their natural environment instead.
The company has already had success with the system. In January 2015, researchers at NovoBiotic reported that they had discovered a novel antibiotic, teixobactin[6]
. “This is a really big win for natural-product drug discovery,” says Ling.
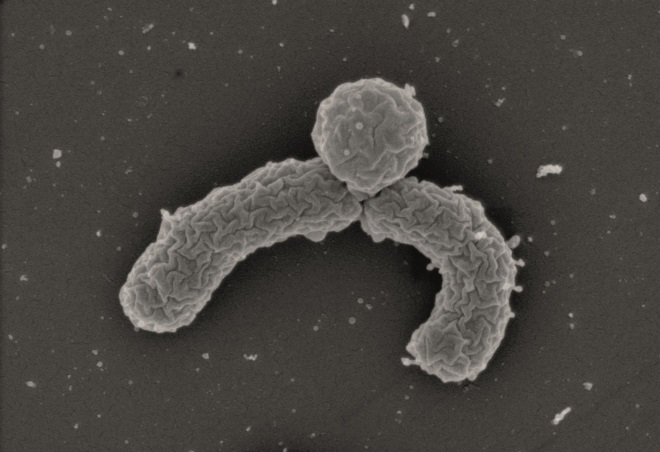
Source: NovoBiotic Pharmaceuticals
A new bacterial strain (pictured) couldn’t previously be grown in the laboratory and has been found to produce the promising antibiotic, teixobactin
“We take bacteria from the environment, mostly from soil, and separate them out individually into chambers with soft agar,” she explains. They are then incubated in the soil, rather than in synthetic media, and form small colonies in each chamber of around 1,000 bacterial cells. “We then pull them back out and find that some will now grow outside their natural environment. We call this domestication.” Ling says that although the discovery of teixobactin is important, she is more excited about the technology that enabled it to happen.
When you look at them under the microscope you can’t believe they all came from nature
NovoBiotic’s technique means that researchers can cultivate an “amazing” variety of bacteria, says Ling. “When you look at them under the microscope you can’t believe they all came from nature.”
The next step in NovoBiotic’s process is to identify which bacteria are completely new, in the hope that they will be able to produce a unique molecule. The researchers take a DNA sample from the individual bacterial colonies and perform genetic analysis, explains Ling. The bacteria’s ribosomal 16S rDNA sequence is compared with the GenBank database of sequenced bacteria to make sure it is unique. The closest match to the 16S rDNA sequence of the bacteria that produced teixobactin was 3% different. By comparison, the DNA from humans and chimpanzees (our closest relatives) diverge by only 1.06%[7]
.
NovoBiotic now has a collection of more than 50,000 bacteria that, according to their genetic profile, were previously uncultivated. “It’s not just about how different a chemical can be, it’s the amazing complexity of structure,” says Ling.
Going to extremes
Researchers at NovoBiotic are currently taking samples only from within the United States, says Ling, because they have not yet exhausted their home turf. But in the quest to find unique microorganisms, other research groups are taking samples from extreme environments, in what is called ‘bioprospecting’. They hope that microbes found in these conditions will have evolved molecules that have never been seen before. This is the strategy adopted by the PharmaSea project, which is funded by the European Union.
“Our philosophy is that nature has done much of the work to evolve molecules that could become drugs,” says Esguerra, who until December 2014 was based at the University ofLeuven, Belgium, and was the PharmaSea project coordinator. The advantage is that these molecules are already designed to target proteins, so then it’s a case of modifying them for human proteins, she explains.
PharmaSea is a collaboration between institutions in eight different European countries and other partners outside Europe. It aims to make natural products attractive to the pharmaceutical industry again, and to identify bottlenecks in this route of drug development. By 2016, when the four-year project ends, the researchers hope to have developed at least two molecules to the point of being ready for clinical trials, one of which should be an antibiotic.
As the name suggests, PharmaSea is scouring the depths of the world’s oceans in a bid to find molecules that the pharmaceutical industry can patent. “We want to tap extreme environments to find novel organisms and therefore hopefully novel secondary metabolites,” explains Esguerra. The average ocean depth is 3,790 metres, and about half of specimens collected from depths greater than 3,000 metres will contain organisms new to science.[8]
In order to go down that deep, PharmaSea has commissioned shipping vessels to take samples from the sea bed. Deep Tek, a UK-based engineering company that provides deep-sea services for the scientific research, salvage, and oil and gas industries, has manufactured something completely novel, says Esguerra: “A 9-km cable drops straight to the sea bed from an ‘opportunity’ vessel.” Samples have already been collected from cold vents in New Zealand and hot vents in the Mariana Trench, the deepest part of the world’s oceans. In spring 2015, Deep Tek plans to take samples from the coast of the Atacama Desert in Chile, the driest place on Earth. “We’re also hoping to go to the Antarctic and Arctic,” says Esguerra.
Once the samples are collected several partner laboratories isolate and catalogue unique microbes. Duplicate samples are sent to each laboratory around Europe to test the bacterial products against a specific target. “Denmark is doing antibiotics, Spain is searching for Alzheimer’s drugs, and others are looking for anticancer drugs,” explains Esguerra, who herself is searching for neuroactive compounds.
Finding novel natural products
Researchers adopt different tactics in pursuit of finding new microbes with the potential to synthesise unique molecules
Above sea level: Researchers are developing new technology that allows them to cultivate microbes that couldn’t previously be grown in the laboratory, at the moment this is estimated to be 99% of bacteria.
Symbiotic bacteria (200 metres below sea level): Many species of sponge possess symbiotic bacteria. A whole new bacterial phylum, Entotheonella, was characterised in 2014 and identified as the producer of almost all the polyketides and peptides associated with the marine sponge host, Theonella swinhoei. This new phylum couldn’t previously be cultivated.
Chemosynthesis (1,000 metres below sea level): As the ocean gets deeper and the light disappears, microbes become dependent on chemosynthesis, rather than photosynthesis, for energy. For researchers these microbes are a source of rich chemical diversity and novel chemical pathways.
New species: Below 3,000m each sample collected is 50% likely to contain a species new to science.
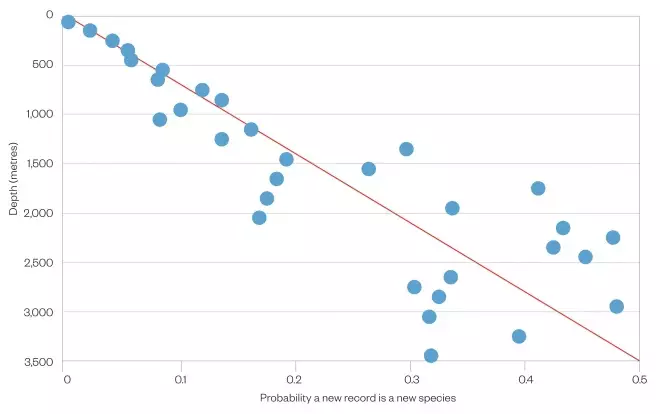
Probability of an ocean specimen being a new species
Source: Carlo Heip / NIOZ / OBIS
Biopiracy
The practice of bioprospecting has sometimes had murky associations with ‘biopiracy’ — the use of a country’s biological resources without the proper consent or compensation of the host country. “You have to make sure that the rights of less developed countries are taken into account and that these nations are not taken advantage of by more industrialised countries,” says Esguerra.
In 1998, a research project called Maya ICBG set out to characterise compounds used by the Maya communities in Mexico. Various US government departments provided funding, with collaboration from Mexican universities and the support of communities and the Mexican government[9]
. But the group was accused of unethical bioprospecting by non-governmental organisations. The criticism centred on the nature of prior informed consent. Eventually the controversy led to the five-year project being terminated after only two years.
The issue of ethics in sourcing compounds is addressed in the Convention on Biological Diversity, which came into force in December 1993 and which 194 states are now party to. One of its main aims is the fair and equitable sharing of the benefits from the utilisation of genetic resources. This goal was formalised in the Nagoya Protocol, which came into force in October 2014.
“In theory, this protocol should promote the utilisation of natural resources by providing more legal certainty,” says PharmaSea collaborator Thomas Vanagt, a marine ecologist who founded the eCOAST marine research centre in Belgium. He led PharmaSea’s input into the Nagoya Protocol.
Vanagt warns that, in practice, the legal situation for researchers and ‘provider’ countries may still not be completely clear. The protocol stipulates that any entity undertaking bioprospecting in a country must abide by that country’s access legislation. “Making access legislation is a national prerogative,” he explains, but not every country has this legislation in place. Costa Rica has had robust legislation for many years, for example, but others still don’t have any.
“Differences in national legislation can cause confusion,” says Vanagt. He believes that one of the biggest issues is that scientists are unaware that their science is regulated. “This legislation will hopefully reduce intentional biopiracy but, importantly, if everyone is aware of the Nagoya Protocol, you won’t have people accidentally falling foul of the law,” he adds.
Access agreements for natural resources are already in place between many countries. Novartis has a number of partners that send it samples in exchange for the transfer of knowledge and technology. However, financial compensation would be expected if a drug sourced from one of these countries was brought to market.
Preserving biodiversity
Thailand’s National Centre for Genetic Engineering and Biotechnology (BIOTEC) is one of Novartis’s partners. The partnership began in 2005 when Thailand’s minister of science and technology, and BIOTEC’s then executive director visited Novartis, says Kanyawim Kirtikara, the current executive director of BIOTEC. “We’ve been screening molecules for a long time but we don’t necessarily have the expertise to bring the compound forward,” she says. This is where Novartis can help.
We’ve been screening molecules for a long time but we don’t necessarily have the expertise to bring the compound forward
It was the richness of Thailand’s biodiversity that first drew BIOTEC to this work. Thailand has a diverse variety of habitats and is home to around 10% of the world’s biodiversity. “Not many microbes have been characterised in Thailand so we want to characterise and preserve Thailand’s biodiversity,” she says. “It’s always a race against the destruction of the habitat. You never know whether the next time you go to a place it will have been bulldozed for a housing development.”
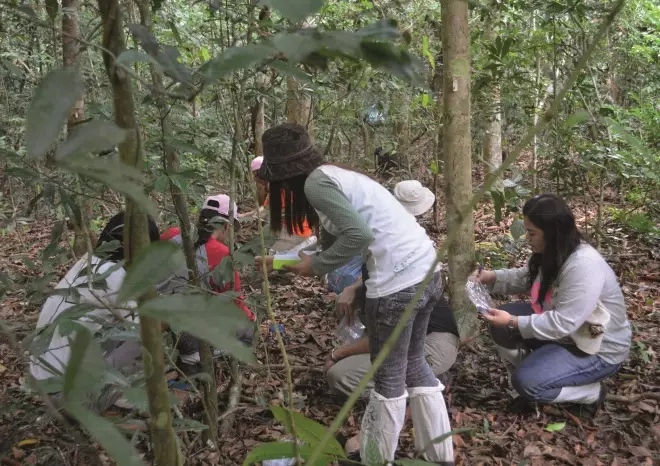

Source: BIOTEC
Scientists from Thailand’s BIOTEC collect samples in the field
Kirtikara says that BIOTEC specialises in collecting fungi that attack insects, and she estimates that it probably has the largest collection of these microbes outside the United States. Scientists at BIOTEC go to a range of habitats within Thailand, such as the limestone region in the centre of the country that has an unusually high pH. Duplicate samples are then sent to Novartis to be evaluated using assays that BIOTEC does not perform.
Frank Petersen, executive director of the natural-products unit at Novartis, says the company has worked hard to set up partnerships with research institutions. Over the past 60 years, Novartis has built up a collection of 70,000 strains of bacteria from many parts of the world, and has a long history of success with natural products. Explaining why Novartis stayed in this line of research when many other pharmaceutical companies jumped ship, Petersen says: “Some said high-throughput screening was the death of natural products, but I saw it as a chance.”
Novartis has miniaturised every step of natural-product discovery, speeding up significantly the deconvolution step from a crude compound mixture to a single substance. “This allows us to identify the active compound in one shot,” says Petersen.
“[Petersen] made the timelines for natural-product drug discovery compatible with our drug development timelines,” says Sylvain Cottens, global head of the Centre for Proteomic Chemistry at Novartis.
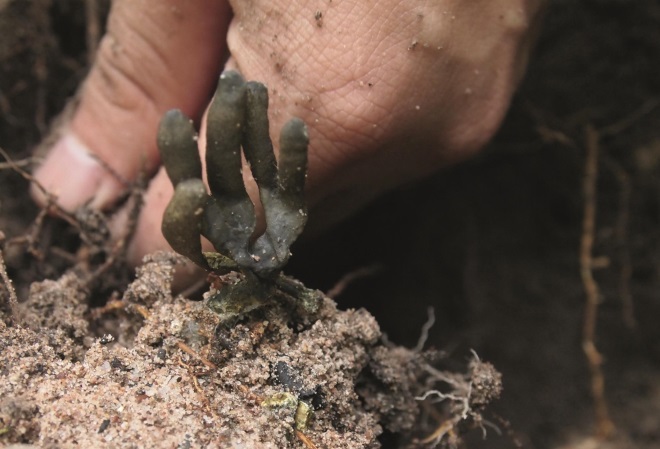

Source: BIOTEC
A scientist from BIOTEC in Thailand, one of Novartis’ partners, collects samples
This is no small feat. After first trying to coax the bacteria into growing, the next hurdle is getting them to produce their full range of molecules. Even then, many different types of bacteria will produce the same set of molecules, which Petersen says has caused a number of companies to leave the field. “But we’ve discovered there are many potential novel compounds that can be found by activating all the different metabolic pathways in the bacteria,” he says. Novartis’s strategy is to systematically switch on every novel secondary metabolic pathway it can find.
Finding the switch
The bacteria are subjected to up to 100 different conditions to try to find the right switch to turn on these pathways. The genes that encode the enzymes to synthesise these metabolites are located next to each other in the bacteria’s genome. Cottens says that they can identify these gene clusters, but occasionally they find one that cannot be activated. They then excise that stretch of DNA and insert it into a ‘super host’ — bacteria that are easy to grow in the lab and have a good ‘precursor flux’, meaning that they have all the metabolic building blocks needed to create these foreign metabolites.
Novartis keeps the metabolites that have a novel chemical structure. “We don’t know the biological function at first. These molecules are added to our screening libraries and tested against all of our drug targets,” says Petersen.
“In the 1990s, a number of companies restricted natural products to just anti-infectives, but Novartis’s success is from screening in all areas,” Petersen explains. Natural products made the field of anti-rejection drugs possible, he says, giving the example from Novartis of everolimus, a derivative of sirolimus, which was found in a soil sample from Easter Island by Brazilian researchers. It is used as an immunosuppressant to prevent organ rejection and to treat kidney cancer.[10]
There is no single right way to find drugs
In the end, Petersen says, the debate around natural products is arbitrary. “There is no single right way to find drugs,” he says. “Drug hits can come from synthetic or natural products and we use both to design and refine molecules.”
Even so, Novartis is unusual in its approach. Despite the interest from academia and small pharmaceutical companies, there is yet to be a large-scale return of ‘big pharma’ to natural-product drug discovery. Ling, from NovoBiotic, isn’t sure there ever will be. “Big pharma don’t always do the right thing scientifically, because they have their bottom-line to think about.”
But Ling believes that if there is enough proof that natural products produce good drug leads, eventually it will be attractive enough for the big companies to return. If they do, it will take “significant investment”, she warns. But she says the more base structures that are discovered, the more ammunition chemists have to hit drug targets. The discovery of teixobactin provides a “whole new backbone to work with and modify”, she says.
Back in Zurich, Piel predicts that the gene-based methods of natural-product research represent a turning point. “We can predict from the gene sequence what many compounds are going to look like without even getting them to be expressed,” he says. His team is hoping to find new enzymes that catalyse new reactions. The next step is biosynthetic engineering, with novel genes being combined with other novel genes to create metabolic pathways not seen in nature, says Piel. “It opens up so many possibilities.”
Going back to nature could be just what the pharmaceutical industry needs.
References
[1] Wilson MC, Mori T, Rückert C et al . An environmental bacterial taxon with a large and distinct metabolic repertoire. Nature 2014; 506:58–62.
[2] Lewis, K. Platforms for antibiotic discovery. Nature Reviews Drug Discovery 2013;12:371–387.
[3] Borchardt JK. The beginnings of drug therapy: Ancient mesopotamian medicine. Drug News Perspective 2002;15:187–192.
[4] Newman, DJ and Cragg GM. Natural products as sources of new drugs over the 30 years from 1981 to 2010. Journal of Natural Products 2012;75: 311−335.
[5] Shetty N and Gupta S. Eribulin drug review. South Asian Journal of Cancer. 2014;3:57–59.
[6] Ling LL, Schneider T, Peoples AJ et al. A new antibiotic kills pathogens without detectable resistance. Nature 2015;517:455–459.
[7] The chimpanzee sequencing and analysis consortium. The initial sequence of the chimpanzee genome and comparison with the human genome. Nature 2005;437:69–87.
[8] Skropeta D. Deep sea natural products. Natural Product Reports 2008;25:1131–1166.
[9] Berlin B and Berlin E. NGOs and the process of prior informed consent in bioprospecting research: the Maya ICBG project in Chiapas, Mexico. International Social Science Journal 2004;55:629–638.
[10] Coppin C. Everolimus: the first approved product for patients with advanced renal cell cancer after sunitinib and/or sorafenib. Biologics 2010;4:91–101.

