Shutterstock.com
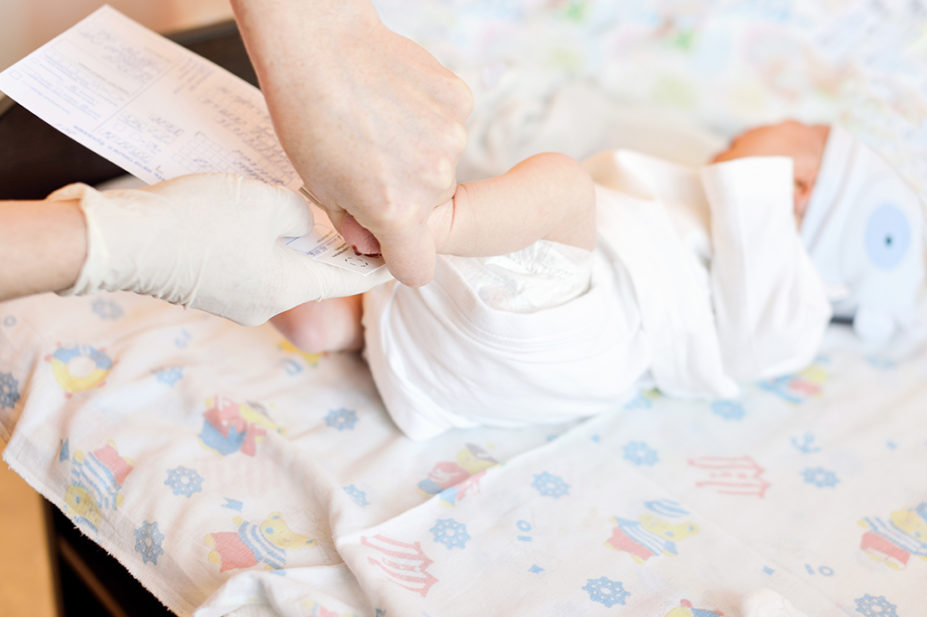
In 2019, then health secretary Matt Hancock announced that he wanted to usher in a genomics revolution by sequencing the genome of every baby in England to map out their risk of disease at birth and offer more personalised healthcare.
While this vision has not materialised in the exact way it was presented back then, it has led to a £105m research project to screen the genomes of 100,000 babies for around 200 rare diseases. On track to begin recruitment in 2023, the plan is to test the waters on the myriad of logistical, financial, ethical and societal challenges that such an approach brings.
England is not alone in this — to date, trials have been relatively small scale, but what was once an abstract concept is now tipping into reality, as teams across the world, including in the United States, Europe, Australia, China and Qatar, grapple with the question of whether genome sequencing could and should be routine (see Box).
At Genomics England — a company set up and owned by the Department of Health and Social Care — advisory committees have been working to identify what diseases should be included in the project, what consent should look like, how samples will be taken and reported, and the pathways of care in place once a rare disease has been identified.
A final list of conditions that will be screened for is yet to be published, but the aim to screen for around 200 diseases is a significant jump from the 9 conditions currently identified through the newborn blood spot test. Yet, that number is relatively conservative compared with programmes in other countries that are looking at reporting up to 500 diseases.
Genomics England stresses that it is only looking at conditions that meet certain criteria, which include: strong evidence that the genetic variance identified causes the condition and can be reliably detected; that a high proportion of children in whom the gene was detected would be expected to have debilitating symptoms; and that early or pre-symptomatic intervention leads to substantially improved outcomes. The condition in question must also affect children by the age of five years.
The conditions screened for will include immune deficiencies, metabolic disorders and rare thyroid conditions. People with immune deficiencies may require a bone marrow transplant but other conditions may be treated with a special diet or simple medicine. One example is biotinidase deficiency, in which the individual is unable to metabolise biotin, which can lead to severe skin conditions, seizures and other neurological disorders, but for which the treatment is a simple supplement. Another is a defect in the THRA gene, which leads to a very rare thyroid resistance condition for which the treatment is thyroxine. Early detection and treatment could help manage conditions but also prevent severe symptoms developing.
David Bick, a clinical geneticist from the United States, who moved to the UK to become clinical advisor for the Newborn Genomes Programme, says: “We’re only going to choose genes we know about. Doctors aren’t going to be doing new treatments that they aren’t already using because this has got to be available to everybody. In the NHS, in many ways, it’s a logistical challenge more than a scientific challenge.”
Around 25–30 trusts will be taking part once the project is fully up and running. Parents-to-be will be taken through the consent processes at antenatal visits so they can weigh up their decision before the baby is born.
Its predecessor, the 100,000 genomes project, set out to look for “actionable results” for rare disease or cancer. Yet, this time, the majority of newborns whose genomes are sequenced will be healthy. If rolled out nationally, Genomics England expects around 3,000 children per year potentially to be identified with one of the diseases screened for, which is about 0.5% of newborn babies. However, in the research phase — which may throw up false positives — they expect to report a finding in around 1–2% of children, who will then be referred for specialist investigation.
Identifying those children with rare inherited conditions is only one part of the research, explains Bick. “Part two is: let’s do some research to identify new conditions and maybe, more importantly, identify treatments for some of the genetic conditions we know about. Then there’s a third part, which is a bit more speculative, which relates to how might a genome be used across a person’s lifespan.”
One group that has been working closely with Genomics England in developing its approach is Genetic Alliance UK, an umbrella organisation of more than 200 patient groups representing rare diseases. These represent families that can speak to the heartache and difficulties faced in getting a diagnosis, which often takes years. This experience of the “diagnostic odyssey” is what proponents of newborn genome screening say they want to avoid.
It’s tremendously powerful to be able to identify conditions where we can make an intervention
Nick Meade, director of policy at Genetic Alliance UK
Nick Meade, director of policy at Genetic Alliance UK, says genome screening has the potential to be very impactful but must be approached with caution. “It’s tremendously powerful to be able to identify conditions where we can make an intervention. We should examine the full potential of this technology but we need to examine that from all sides and make sure that we’re managing any risks that come with it.”
What often gets missed, says Meade, is that the current newborn heel prick test could already be expanded to look at more conditions, which he would like to see addressed in parallel. “The UK falls behind the majority of European countries in terms of how many conditions we’re screening for, there are 9 conditions that 15 countries in Europe are screening for that we’re not.”
Given the large unmet health need for children with rare conditions, and how devastating they can be, the general ethos among families is “leave no stone unturned”, says Meade. “But we don’t want to go too far beyond what the public is comfortable with because you can damage public perception of technology in a way that can undermine the progress that we’ve made. The concern from our community would be making sure that the whole population of the UK is well enough informed and supports what we’re doing and that [the genomic data] is stored safely [and] securely, with adequate privacy.”
As with the 100,000 genomes project, the genomic data will be added to a secure database to which university researchers, life sciences or pharmaceutical companies have to apply for access, giving details of the work they intend to do. The genetic information, coupled with clinical data from healthcare records, will be available in the National Genomics Research Library, where any analysis has to be done within the cloud-based system, with only summary data available for download.
The Pharmaceutical Journal has learnt, via a Freedom of Information request, that since 2019, 78 commercial applications and 451 academic and clinical researchers have had their applications to access the research database approved, with no cases rejected. Industry partners listed on the Genomics England website include some of the world’s biggest pharmaceutical companies, such as AstraZeneca, GSK, Boerhinger Ingelheim, Novartis, Roche, Takeda and MSD.
David Curtis, honorary professor at the University College London Genetics Institute, has strong concerns about the whole approach to newborn genome screening, and believes it is hard to justify ethically in a healthy population who cannot themselves consent.
The thing that tends to get very understated is how difficult it is to interpret results from genome sequencing
David Curtis, honorary professor at the University College London Genetics Institute
“Once in a very, very, blue moon, you will pick something up — they are very tiny numbers. The thing that tends to get very understated is how difficult it is to interpret results from genome sequencing,” he adds. Even when people are going into a research project with a disease to try to find a diagnosis, it’s unusual to get a definitive answer, he explains.
The value of the data, says Curtis, is therefore not necessarily for the individual. “It’s being sold to research institutions, universities and biopharma companies, and they will pay for that. When you’re signing up to get your genome sequenced, you’re also signing up to allow these other companies and collaborators and universities to have ongoing access to your health outcomes.”
He is also not overly convinced of how useful genome sequences are from a scientific standpoint. “I’ve analysed a lot of UK Biobank [a large biomedical database containing in-depth genetic and health information from half a million UK participants] and it’s surprising how little you can get out. In the context of a national health service, where you need nurses, you can’t get ambulances, you can’t get doctors, this is not a good use of resources.”
Some may feel he overstates the risks of abuse of genomic data because safeguards are put in place, Curtis muses. “At the moment, it’s illegal to abuse the data but what will happen in ten years’ time? This government has trampled over all sorts of things we thought were fairly solid. I think it’s legitimate to factor that in when you’re giving away data of newborn babies.”
The cost-effectiveness, acceptability of screening among parents and the health outcomes of children who are screened will all be part of the study. The National Screening Committee, who have had representation in the working groups putting the research project together, will ultimately have to decide whether to proceed on a larger scale. Those leading the study say that what is acceptable to parents in the UK may be entirely different to what is acceptable to those in other parts of the world.
Rachel Horton, clinical research fellow at the Wellcome Centre for Human Genetics in Oxford, has written about several challenges that come with the ability to sequence genomes easily and quickly. Newborn programmes “must tread a delicate balance” between identifying as many babies with serious but treatable conditions as possible, yet not raising unjustified anxiety around healthy babies, she says. Factors, such as the risk of false positives and uncertain findings that lead to a lifetime of tests, are often missing from public discussions — the initial announcement from Matt Hancock, and the subsequent coverage in the media, does not reflect the difficult decisions that must be made in reality, she notes.
“My impression is that [Genomics England] are going to take a very conservative stance as to what they report from genome sequencing. [But] whether you could answer similar questions [by collecting] less data is a big question. [And] it also introduces potential uncertainties into the health service around [whether it is] set up to deal with this and what happens to those who already have symptomatic disease.”
In some cases, results may not be as clear cut as the general public may think, she explains.
“I think it’s really important that the programme is raising those issues with parents in the consent process and also, with the volume of data they’re asking for, what portion is going to be used for research versus what’s going to be looked at for the child’s direct benefit.”
In the long run — if approached in the right way — Horton believes that collecting this data will have benefit for society. What is vital, she says, is that those signing up to the programme fully understand what the goal is. “All screening has a cost and what the cost-benefit ratio is going to be for babies is not a straightforward question.”
Newborn screening programmes around the world
Europe
Screen4Care
Funded by the EU, academia and the pharmaceutical industry, Screen4Care has 36 partners across 15 countries. In a pilot study, it wants to use whole-genome sequencing in at least 18,000 newborns to look for treatable and actionable diseases. It is also looking at algorithms to detect early symptoms in infant health records.
Australia
Baby Beyond
Baby Beyond is one of three projects underway in Australia, after the government announced a tranche of funding. This has grown from earlier research looking specifically at newborn hearing loss. Now the team plan to use whole-genome sequencing in 1,000 newborns to look for up to 500 conditions, but with the final list yet to be agreed.
United States
BabySeq
First set up in 2013, this randomised controlled trial, at Brigham and Women’s Hospital, Boston, and Boston Children’s Hospital, of 325 newborns — of which 9% had a risk variant for a childhood onset disease and 2% for an adult-onset condition. Now, in phase two, 1,000 babies from diverse populations will be screened using whole-genome analysis.
The GUARDIAN project
Led by a team at Columbia University in New York, this project, which stands for Genomic Uniform-screening Against Rare Diseases in All Newborns, hopes to enrol 100,000 newborns over four years, initially looking at 250 conditions. So far, uptake has reached 70% with 250 babies enrolled, the researchers have said.
ScreenPlus
Another New York team, this time at Albert Einstein College of Medicine and the Children’s Hospital at Montefiore, is focusing on biochemical testing in 175,000 babies, but is using confirmatory disease-gene sequencing and may collaborate with genomics programmes at some point.
Early Check
In North Carolina, a team of researchers began screening for spinal muscular atrophy and fragile X syndrome four years ago. Now, a new sequencing project set to start in 2023 will analyse the genomes of 10,000 newborns for 200 childhood-onset monogenic conditions and follow children for 23 months.
BeginNGS
A consortium led by Rady Children’s Institute for Genomic Medicine in California aims to provide a platform for implementing whole-genome sequencing and connect doctors with the interventions needed, as well as help rare disease drug development. A small pilot study has been done and there are now plans to screen 2,000 infants at sites across the United States and Greece for up to 500 disorders.