Sinclair Stammers / Science Photo Library
The genomes of malaria parasites contain many genes of unknown function. However, knowing the genes and pathways that contribute to parasite growth is critical to guiding the discovery of new drug targets.
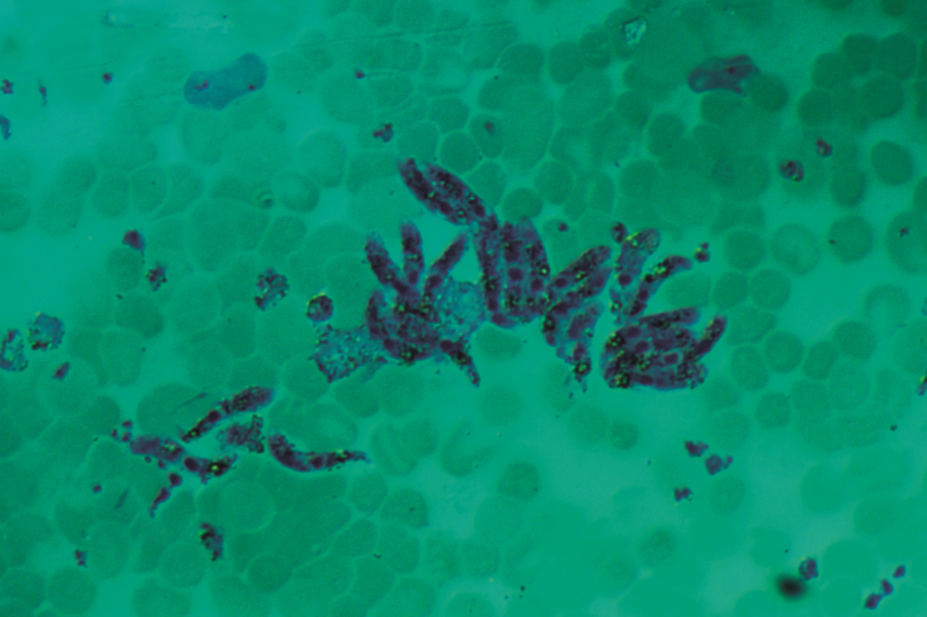
In the first large-scale study of malaria gene function, researchers at the Wellcome Trust Sanger Institute analysed more than half of the genes in the genome of one species of malaria parasite, Plasmodium berghei. To do this, they measured the growth rates in mice of 2,578 P. berghei knockout mutants, each of which were tagged with a unique barcode.
It was found that during a single blood stage of its life cycle, the P. berghei parasite requires around two-thirds of the genes looked at to develop normally.
Publishing their results in Cell
[1]
(13 July), the researchers say that this shows there are many more potential targets for new antimalarial drug development than previously thought.
References
[1] Bushell E, Gomes A, Sanderson T et al. Functional profiling of a Plasmodium genome reveals an abundance of essential genes. Cell 2017;170:260–272. doi: 10.1016/j.cell.2017.06.030