Phototake Inc. / Alamy
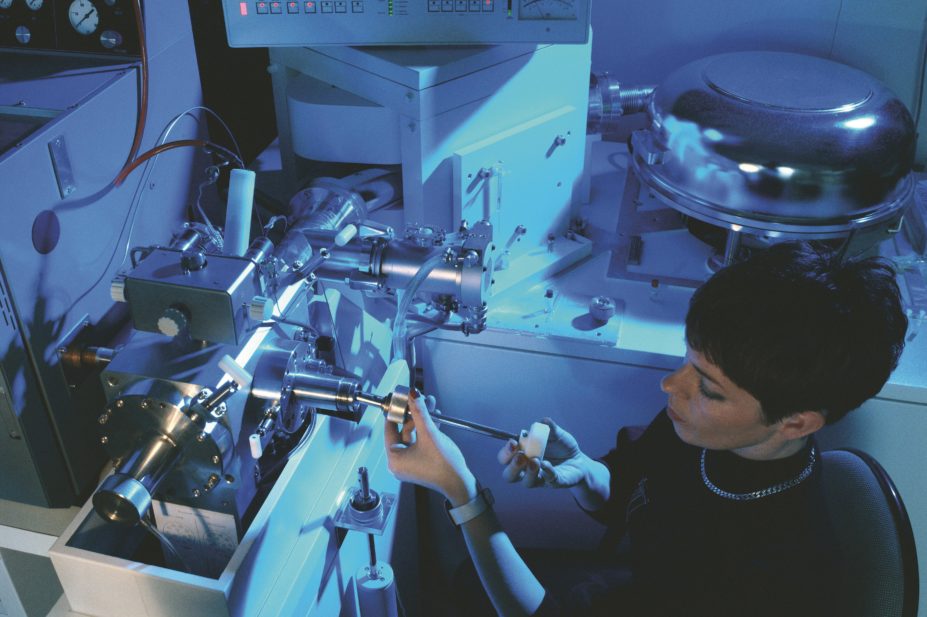
A mass spectroscopy technique able to identify bacterial infections within minutes could lead to quicker administering of the correct treatment, a study presented at the American Society for Microbiology’s annual ICAAC meeting in Washington, DC, found.
The technique can accurately identify a microbe within two to three minutes, according to Kathryn French of the Heart of England NHS Foundation Trust, in Birmingham, while traditional culture analysis can take up to 48 hours.
“The technique has been well-established and is widely-used on plate culture,” said French, one of the study authors.
In the study[1]
, the process – known as matrix-assisted laser desorption/ionization time of flight mass spectroscopy (MALDI-ToF) – came up with three discordant results out of 115. The technique allowed for positive identification of a given microbe in 65% of cases where the infection was Gram positive, and 85% of cases where it was Gram negative, French added.
In the study, researchers retrospectively reviewed data from all patients at the hospital with blood cultures whose Gram stains were positive for microorganisms.
They compared the clinical advice given on the first day (with only a Gram stain result) with the follow-up advice given on day two (after the sample had been conventionally cultured). The outcomes were also compared to what the MALDI-ToF technique would have found.
In 28 of the 115 cases (24.3%), using the MALDI-ToF technique immediately would have had a clear clinical benefit. In 11 cases (9.6%), identification of the correct microorganism would have led to a different prescription for antibiotics, meaning that five patients would have received the correct antibiotic 24 hours sooner.
A further 14 cases (12%) would have been identified as clinically insignificant and unnecessary communications with medical staff could have been avoided.
However, in some cases, the MALDI-ToF technique was unable to detect the nature of the infectious organism and would have yielded no benefit. “If the Gram scan on day one was mixed, because there were two organisms in there, the device struggles to differentiate the two, and can’t come up with the identification,” French explained.
References
[1] French KE, Evans J, Tanner H et al. Direct identification of bacteria from positive blood cultures using MALDI-ToF - the clinical impact. [Poster D-903] American Society for Microbiology. Interscience Conference on Antimicrobial Agents and Chemotherapy. Washington DC. 7 September 2014.